X-ray screening identifies potential candidates for corona drugs
With the help of DESY's X-ray light source PETRA III, a research team has identified several candidates for possible active substances that bind to an important protein of the coronavirus SARS-CoV-2 and could thus be a basis for a drug against the infection. In a high-throughput procedure, the researchers, under the leadership of DESY and the University of Hamburg, partly from CFEL, are currently testing more than 5,600 known active substances from a library of the Fraunhofer Institute for Molecular Biology in a so-called X-ray screening. After measuring a total of 5,768 samples containing 3893 different active substances, the scientists have so far been able to identify a total of 13 substances that bind to a viral protein. The next steps will now be to test whether these substances inhibit protein activity and slow down the multiplication of the virus.
The robot arm allows a fast, automatic change of the frozen samples (middle). Credit: DESY, Marta Mayer
An important starting point in the search for an effective drug against the coronavirus SARS-CoV-2 is to stop its multiplication in the host's body: Viruses cannot replicate on their own. To do this, they hijack cells of their host, introduce their own genetic material into the cells and induce them to produce new viruses. Proteins play an important role in all these steps. If it is possible to block a key protein, it may be possible to interrupt replication and thus defeat the infection.
The P11 beamline of DESY's PETRA III research light source specializes in structural biological investigations. Here, the three-dimensional spatial structure of proteins can be resolved with atomic precision. The research team used this to examine several thousand active substances already in existence for the treatment of other diseases to find out whether and how they “dock” to one of the key proteins responsible for the reproduction of the virus. With this investigation, the team hopes to shorten the path to a drug: While the development of an approved drug from a new compound usually takes several years, some of the substances investigated in this study have already been approved for the treatment of humans or are at least in the trial stage.
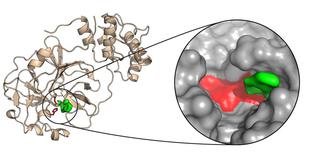
The main protease of SARS-CoV-2 is a crucial component in the replication of the virus. It carries out its work in the so-called active center (shown here in red). X-ray examinations have now revealed a substance (green) that binds firmly to the active center. Credit: DESY, Sebastian Günther
In a high-throughput process, the researchers were able to test around 70 percent of the various active ingredients in two weeks. Thanks to a fully automatic sample change with a robotic arm, each measurement took only three minutes. “With the help of automated data analysis, we were able to identify 13 active substances that bind to the proteins,” says DESY researcher Alke Meents. “One of them binds covalently, that is a very strong chemical bond, at a central location to the main protease of the virus, making it a particularly promising candidate.” The main protease is a key protein for the reproduction of the virus.
Proof that the active ingredient binds to the viral multiplication protein is the first step on the way to a drug. “Next, researchers at the Universities of Lübeck and Hamburg and the Fraunhofer Institute will now investigate in the laboratory whether this active substance also inhibits protein activity,” says virus researcher Rolf Hilgenfeld from the University of Lübeck, whose research group is involved in the screening. In a third step, the Hamburg-based Bernhard Nocht Institute plans to test in cell cultures whether the substance inhibits or even prevents the virus from multiplying.
Researchers from the universities of Hamburg and Lübeck, the Fraunhofer Institute for Molecular Biology, the Bernhard Nocht Institute for Tropical Medicine, the European Molecular Biology Laboratory, the Max Planck Society, Helmholtz-Zentrum Berlin and DESY are involved in the work.